|
Doxygen attempts to fulfill that same gap, but it also suffers from trying to support every language under the sun, as well as the inherent issues of C/C++ (the use of the preprocessor to generate module boundaries does not make for fun processing). Similarly, Rust also has a widely-used builtin documentation tool (and again, the official standard library documentation is entirely the result of rustdoc).Ĭ/C++ of course don't have a builtin tool. If you've ever used, you've used the results of Javadoc. Javadoc was the first major such tool if I'm trying to use a Java library, I don't read the source code, I start by reading the generated documentation. 
If you look at other languages, automatically-generated code documentation has become the main way people ingest documentation for software projects. The problem isn't that no one reads generated codedoc, the problem is that doxygen is a pretty abysmal code generator. "Installing Doxygen on a project" achieves nothing, one has to write the actual docs first. So here I threw away all this noise and the theme is actively forcing the library authors to focus on important stuff in the docs, explicitly excluding useless things that could be auto-generated. 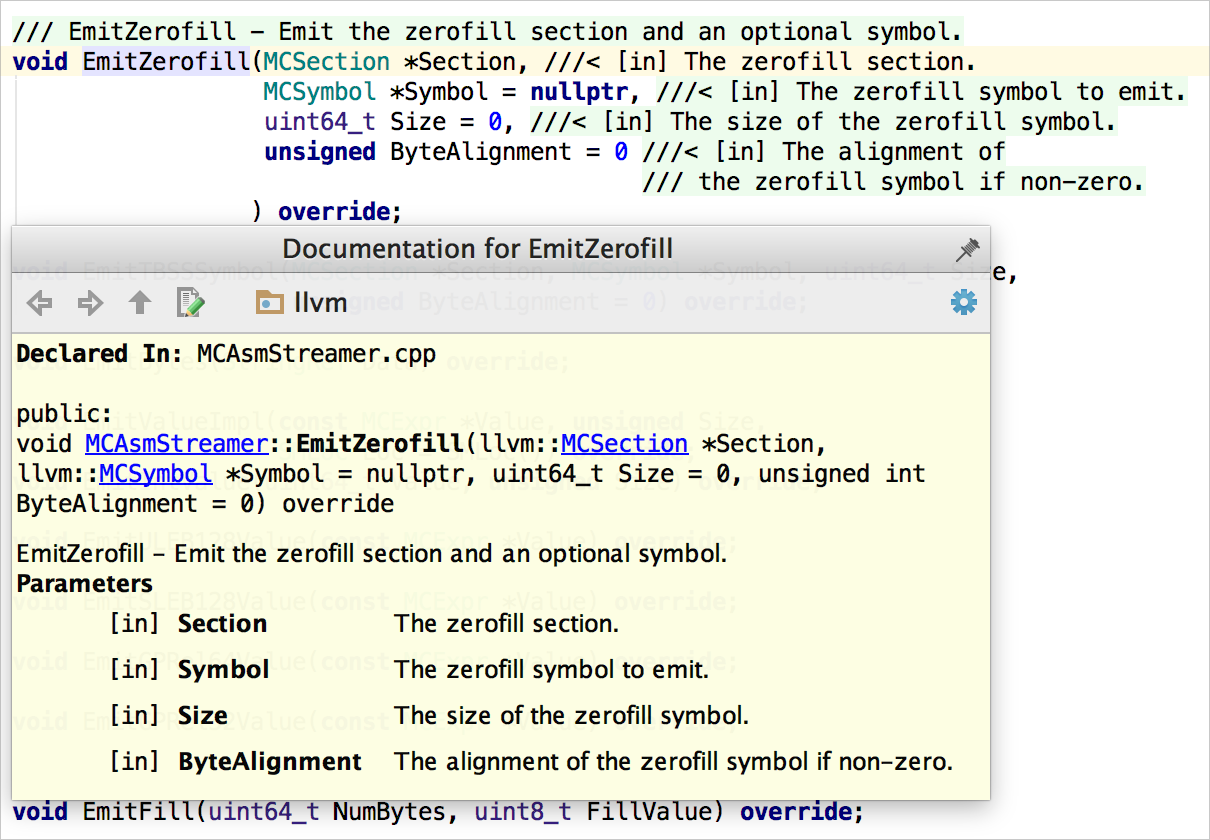
Not to see stuff that's already explained by the code itself. The user should visit docs to get to know a high-level overview of a library or human-readable explanation of an algorithm.  
Doxygen implicitly generates useless info about what file is included by what file, list of places from where a function gets called, what line was this and that function declaration in (and where is the definition), huge class inheritance diagrams, entangled monstrous file dependency diagrams, alphabetical file index, alphabetical symbol index, including every possible undocumented symbol and file it can find and tons and tons of other stuff that has a total value of 0. Library ( dplyr ) starwars %>% filter ( species = "Droid" ) #> # A tibble: 6 × 14 #> name height mass hair_color skin_color eye_color birth_year sex gender #> #> 1 C-3PO 167 75 gold yellow 112 none masculi… #> 2 R2-D2 96 32 white, blue red 33 none masculi… #> 3 R5-D4 97 32 white, red red NA none masculi… #> 4 IG-88 200 140 none metal red 15 none masculi… #> 5 R4-P17 96 NA none silver, red red, blue NA none feminine #> # ℹ 1 more row #> # ℹ 5 more variables: homeworld, species, films, #> # vehicles, starships starwars %>% select ( name, ends_with ( "color" ) ) #> # A tibble: 87 × 4 #> name hair_color skin_color eye_color #> #> 1 Luke Skywalker blond fair blue #> 2 C-3PO gold yellow #> 3 R2-D2 white, blue red #> 4 Darth Vader none white yellow #> 5 Leia Organa brown light brown #> # ℹ 82 more rows starwars %>% mutate ( name, bmi = mass / ( ( height / 100 ) ^ 2 ) ) %>% select ( name : mass, bmi ) #> # A tibble: 87 × 4 #> name height mass bmi #> #> 1 Luke Skywalker 172 77 26.0 #> 2 C-3PO 167 75 26.9 #> 3 R2-D2 96 32 34.7 #> 4 Darth Vader 202 136 33.3 #> 5 Leia Organa 150 49 21.8 #> # ℹ 82 more rows starwars %>% arrange ( desc ( mass ) ) #> # A tibble: 87 × 14 #> name height mass hair_color skin_color eye_color birth_year sex gender #> #> 1 Jabba De… 175 1358 green-tan… orange 600 herm… mascu… #> 2 Grievous 216 159 none brown, wh… green, y… NA male mascu… #> 3 IG-88 200 140 none metal red 15 none mascu… #> 4 Darth Va… 202 136 none white yellow 41.9 male mascu… #> 5 Tarfful 234 136 brown brown blue NA male mascu… #> # ℹ 82 more rows #> # ℹ 5 more variables: homeworld, species, films, #> # vehicles, starships starwars %>% group_by ( species ) %>% summarise ( n = n ( ), mass = mean ( mass, na.rm = TRUE ) ) %>% filter ( n > 1, mass > 50 ) #> # A tibble: 8 × 3 #> species n mass #> #> 1 Droid 6 69.8 #> 2 Gungan 3 74 #> 3 Human 35 82.8 #> 4 Kaminoan 2 88 #> 5 Mirialan 2 53.Exactly.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
